
PostgreSQL is emerging as the standard destination for database migrations from proprietary databases. As a consequence, there is an increase in demand for database side code migration and associated performance troubleshooting. One might be able to trace the latency to a plsql function, but explaining what happens within a function could be a difficult question. Things get messier when you know the function call is taking time, but within that function there are calls to other functions as part of its body. It is a very challenging question to identify which line inside a function—or block of code—is causing the slowness. In order to answer such questions, we need to know how much time an execution spends on each line or block of code. The plprofiler project provides great tooling and extensions to address such questions.
Demonstration of plprofiler using an example
The plprofiler source contains a sample for testing plprofiler. This sample serves two purposes. It can be used for testing the configuration of plprofiler, and it is great place to see how to do the profiling of a nested function call. Files related to this can be located inside the “examples” directory. Don’t worry—I’ll be running through the installation of plprofiler later in this article.
$ cd examples/
The example expects you to create a database with name “pgbench_plprofiler”
postgres=# CREATE DATABASE pgbench_plprofiler; CREATE DATABASE
The project provides a shell script along with a source tree to test plprofiler functionality. So testing is just a matter of running the shell script.
$ ./prepdb.sh dropping old tables... ....
Running session level profiling
This profiling uses session level local-data. By default the plprofiler extension collects runtime data in per-backend hashtables (in-memory). This data is only accessible in the current session, and is lost when the session ends or the hash tables are explicitly reset. plprofiler’s run command will execute the plsql code and capture the profile information.
This is illustrated by below example,
$ plprofiler run --command "SELECT tpcb(1, 2, 3, -42)" -d pgbench_plprofiler --output tpcb-test1.html SELECT tpcb(1, 2, 3, -42) -- row1: tpcb: -42 ---- (1 rows) SELECT 1 (0.073 seconds)
What happens during above plprofiler command run can be summarised in 3 steps:
- A function call with four parameters “SELECT tpcb(1, 2, 3, -42)” is presented to the plprofiler tool for execution.
- plprofiler establishes a connection to PostgreSQL and executes the function
- The tool collects the profile information captured in the local-data hash tables and generates an HTML report “tpcb-test1.html”
Global profiling
As mentioned previously, this method is useful if we want to profile the function executions in other sessions or on the entire database. During global profiling, data is captured into a shared-data hash table which is accessible for all sessions in the database. The plprofiler extension periodically copies the local-data from the individual sessions into shared hash tables, to make the statistics available to other sessions. See the
plprofiler monitor
command, below, for details. This data still relies on the local database system catalog to resolve Oid values into object definitions.
In this example, the plprofiler tool will be running in monitor mode for a duration of 60 seconds. Every 10 seconds, the tool copies data from local-data to shared-data.
$ plprofiler monitor --interval=10 --duration=60 -d pgbench_plprofiler monitoring for 60 seconds ... done.
For testing purposes you can start executing a few functions at the same time.
Once the data is captured into shared-data, we can generate a report. For example:
$ plprofiler report --from-shared --title=MultipgMax --output=MultipgMax.html -d pgbench_plprofiler
The data in shared-data will be retained until it’s explicitly cleared using the
plprofiler reset
command
$ plprofiler reset
If there is no profile data present in the shared hash tables, execution of the report will result in error message.
$ plprofiler report --from-shared --title=MultipgMax --output=MultipgMax.html
Traceback (most recent call last):
File "/usr/bin/plprofiler", line 11, in <module>
load_entry_point('plprofiler==4.dev0', 'console_scripts', 'plprofiler')()
File "/usr/lib/python2.7/site-packages/plprofiler-4.dev0-py2.7.egg/plprofiler/plprofiler_tool.py", line 67, in main
return report_command(sys.argv[2:])
File "/usr/lib/python2.7/site-packages/plprofiler-4.dev0-py2.7.egg/plprofiler/plprofiler_tool.py", line 493, in report_command
report_data = plp.get_shared_report_data(opt_name, opt_top, args)
File "/usr/lib/python2.7/site-packages/plprofiler-4.dev0-py2.7.egg/plprofiler/plprofiler.py", line 555, in get_shared_report_data
raise Exception("No profiling data found")
Exception: No profiling data foundReport on profile information
The HTML report generated by plprofiler is a self-contained HTML document and it gives detailed information about the PL/pgSQL function execution. There will be a clickable FlameGraph at the top of the report with details about functions in the profile. The plprofiler FlameGraph is based on the actual Wall-Clock time spent in the PL/pgSQL functions. By default, plprofiler provides details on the top ten functions, based on their self_time (total_time – children_time).

This section of the report is followed by tabular representation of function calls. For example:
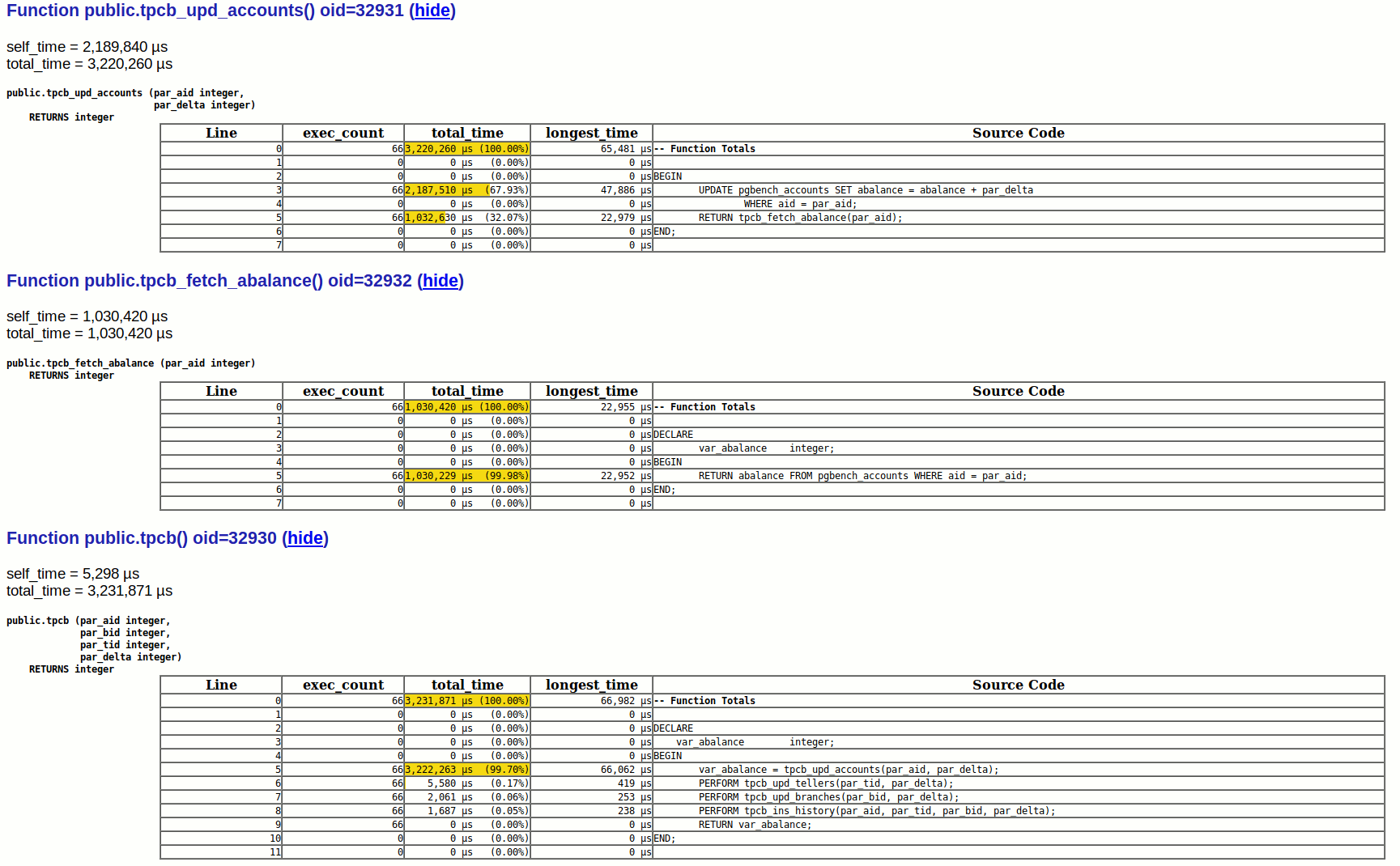
This gives a lot of detailed information such as execution counts and time spend against each line of code.
Binary Packages
Binary distributions of plprofiler are not common. However the BigSQL project provides plprofiler packages as an easy to use bundle. Such ready-to-use packages are one of the reasons for BigSQL to remain as one of the most developer friendly PostgreSQL distributions. The first screen of Package manager installation of BigSQL provided me with the information I am looking for:

Appears that there was a recent release of BigSQL packages and plprofiler is an updated package within that.
Installation and configuration is made simple:
$ ./pgc install plprofiler-pg11 ['plprofiler-pg11'] File is already downloaded. Unpacking plprofiler-pg11-3.3-1-linux64.tar.bz2 install-plprofiler-pg11... Updating postgresql.conf file: old: #shared_preload_libraries = '' # (change requires restart) new: shared_preload_libraries = 'plprofiler'
As we can see, even PostgreSQL parameters are updated to have plprofiler as a
shared_preload_library
. If need to use plprofiler for investigating code, these binary packages from the BigSQL project are my first preference because everything is ready to use. Definitely, this is developer-friendly.
Creation of extension and configuring the plprofiler tool
At the database level, we should create the plprofiler extension to profile the function execution. This step needs to be performed in both cases, whether we want global profiling where share_preload_libraries are set, or at session level where that is not required
postgres=# create extension plprofiler; CREATE EXTENSION
plprofiler is not just an extension, but comes with tooling to invoke profiling or to generate reports. These scripts are primarily coded in Python and uses psycopg2 to connect to PostgreSQL. The python code is located inside the “python-plprofiler” directory of the source tree. There are a few perl dependencies too which will be resolved as part of installation
sudo yum install python-setuptools.noarch sudo yum install python-psycopg2 cd python-plprofiler/ sudo python ./setup.py install
Building from source
If you already have a PostgreSQL instance running using binaries from PGDG repository OR you want to wet your hands by building everything from source, then installation needs a different approach. I have PostgreSQL 11 already running on the system. The first step is to get the corresponding development packages which have all the header files and libraries to support a build from source. Obviously this is the thorough way of getting plprofiler working.
$ sudo yum install postgresql11-devel
We need to have build tools, and since the core of plprofiler is C code, we have to install a C compiler and make utility.
$ sudo yum install gcc make
Preferably, we should build plprofiler using the same OS user that runs PostgreSQL server, which is “postgres” in most environments. Please make sure that all PostgreSQL binaries are available in the path and that you are able to execute the pg_config, which lists out build related information:
$ pg_config BINDIR = /usr/pgsql-11/bin .. INCLUDEDIR = /usr/pgsql-11/include PKGINCLUDEDIR = /usr/pgsql-11/include INCLUDEDIR-SERVER = /usr/pgsql-11/include/server LIBDIR = /usr/pgsql-11/lib PKGLIBDIR = /usr/pgsql-11/lib LOCALEDIR = /usr/pgsql-11/share/locale MANDIR = /usr/pgsql-11/share/man SHAREDIR = /usr/pgsql-11/share SYSCONFDIR = /etc/sysconfig/pgsql PGXS = /usr/pgsql-11/lib/pgxs/src/makefiles/pgxs.mk CONFIGURE = '--enable-rpath' '--prefix=/usr/pgsql-11' '--includedir=/usr/pgsql-11/include' '--mandir=/usr/pgsql-11/share/man' '--datadir=/usr/pgsql-11/share' '--with-icu' 'CLANG=/opt/rh/llvm-toolset-7/root/usr/bin/clang' 'LLVM_CONFIG=/usr/lib64/llvm5.0/bin/llvm-config' '--with-llvm' '--with-perl' '--with-python' '--with-tcl' '--with-tclconfig=/usr/lib64' '--with-openssl' '--with-pam' '--with-gssapi' '--with-includes=/usr/include' '--with-libraries=/usr/lib64' '--enable-nls' '--enable-dtrace' '--with-uuid=e2fs' '--with-libxml' '--with-libxslt' '--with-ldap' '--with-selinux' '--with-systemd' '--with-system-tzdata=/usr/share/zoneinfo' '--sysconfdir=/etc/sysconfig/pgsql' '--docdir=/usr/pgsql-11/doc' '--htmldir=/usr/pgsql-11/doc/html' 'CFLAGS=-O2 -g -pipe -Wall -Wp,-D_FORTIFY_SOURCE=2 -fexceptions -fstack-protector-strong --param=ssp-buffer-size=4 -grecord-gcc-switches -m64 -mtune=generic' 'LDFLAGS=-Wl,--as-needed' 'PKG_CONFIG_PATH=:/usr/lib64/pkgconfig:/usr/share/pkgconfig' CC = gcc ... VERSION = PostgreSQL 11.1
Now we’re ready to get the source code and build it. You should be able to checkout the git repository for plprofiler.
$ git clone https://github.com/pgcentral/plprofiler.git Cloning into 'plprofiler'... ...
Building against PostgreSQL 11 binaries from PGDG can be a bit complicated because of th JIT feature. Configuration flag
--with-llvm
will be enabled. So we may have to have LLVM present in the system as detailed in my previous blog about JIT in PostgreSQL11
Once we’re ready, we can move to the plprofiler directory and build it:
$ cd plprofiler $ make USE_PGXS=1 --- Output ---- gcc -Wall -Wmissing-prototypes -Wpointer-arith -Wdeclaration-after-statement -Wendif-labels -Wmissing-format-attribute -Wformat-security -fno-strict-aliasing -fwrapv -fexcess-precision=standard -O2 -g -pipe -Wall -Wp,-D_FORTIFY_SOURCE=2 -fexceptions -fstack-protector-strong --param=ssp-buffer-size=4 -grecord-gcc-switches -m64 -mtune=generic -fPIC -I. -I./ -I/usr/pgsql-11/include/server -I/usr/pgsql-11/include/internal -D_GNU_SOURCE -I/usr/include/libxml2 -I/usr/include -c -o plprofiler.o plprofiler.c gcc -Wall -Wmissing-prototypes -Wpointer-arith -Wdeclaration-after-statement -Wendif-labels -Wmissing-format-attribute -Wformat-security -fno-strict-aliasing -fwrapv -fexcess-precision=standard -O2 -g -pipe -Wall -Wp,-D_FORTIFY_SOURCE=2 -fexceptions -fstack-protector-strong --param=ssp-buffer-size=4 -grecord-gcc-switches -m64 -mtune=generic -fPIC -shared -o plprofiler.so plprofiler.o -L/usr/pgsql-11/lib -Wl,--as-needed -L/usr/lib64/llvm5.0/lib -L/usr/lib64 -Wl,--as-needed -Wl,-rpath,'/usr/pgsql-11/lib',--enable-new-dtags /opt/rh/llvm-toolset-7/root/usr/bin/clang -Wno-ignored-attributes -fno-strict-aliasing -fwrapv -O2 -I. -I./ -I/usr/pgsql-11/include/server -I/usr/pgsql-11/include/internal -D_GNU_SOURCE -I/usr/include/libxml2 -I/usr/include -flto=thin -emit-llvm -c -o plprofiler.bc plprofiler.c
Now we should be able to install this extension:
$ sudo make USE_PGXS=1 install --- Output ---- /usr/bin/mkdir -p '/usr/pgsql-11/lib' /usr/bin/mkdir -p '/usr/pgsql-11/share/extension' /usr/bin/mkdir -p '/usr/pgsql-11/share/extension' /usr/bin/install -c -m 755 plprofiler.so '/usr/pgsql-11/lib/plprofiler.so' /usr/bin/install -c -m 644 .//plprofiler.control '/usr/pgsql-11/share/extension/' /usr/bin/install -c -m 644 .//plprofiler--1.0--2.0.sql .//plprofiler--2.0--3.0.sql .//plprofiler--3.0.sql '/usr/pgsql-11/share/extension/' /usr/bin/mkdir -p '/usr/pgsql-11/lib/bitcode/plprofiler' /usr/bin/mkdir -p '/usr/pgsql-11/lib/bitcode'/plprofiler/ /usr/bin/install -c -m 644 plprofiler.bc '/usr/pgsql-11/lib/bitcode'/plprofiler/./
The above command expects all build tools to be in the proper path even with sudo.
Profiling external sessions
To profile a function executed by another session, or all other sessions, we should load the libraries at global level. In production environments, that will be the case. This can be done by adding the extension library to the
shared_preload_libraries
specification. You won’t need this if you only want to profile functions executed within your session. Session level profiling is generally possible only in Dev/Test environments.
To enable global profiling, verify the current value of
shared_preload_libraries
and add plprofiler to the list.
postgres=# show shared_preload_libraries ; shared_preload_libraries -------------------------- (1 row) postgres=# alter system set shared_preload_libraries = 'plprofiler'; ALTER SYSTEM postgres=#
This change requires us to restart the PostgreSQL server
$ sudo systemctl restart postgresql-11
After the restart, it’s a good idea to verify the parameter change
postgres=# show shared_preload_libraries ; shared_preload_libraries -------------------------- plprofiler (1 row)
From this point onwards, the steps are same as those for the binary package setup discussed above.
Summary
plprofiler is a wonderful tool for developers. I keep seeing many users who are in real need of it. Hopefully this blog post will help those who never tried it.

 Modern CPU models have a huge number of cores. For many years, applications have been sending queries in parallel to databases. Where there are reporting queries that deal with many table rows, the ability for a query to use multiple CPUs helps us with a faster execution. Parallel queries in PostgreSQL allow us to utilize many CPUs to finish report queries faster. The parallel queries feature was implemented in 9.6 and helps. Starting from PostgreSQL 9.6 a report query is able to use many CPUs and finish faster.
Modern CPU models have a huge number of cores. For many years, applications have been sending queries in parallel to databases. Where there are reporting queries that deal with many table rows, the ability for a query to use multiple CPUs helps us with a faster execution. Parallel queries in PostgreSQL allow us to utilize many CPUs to finish report queries faster. The parallel queries feature was implemented in 9.6 and helps. Starting from PostgreSQL 9.6 a report query is able to use many CPUs and finish faster.